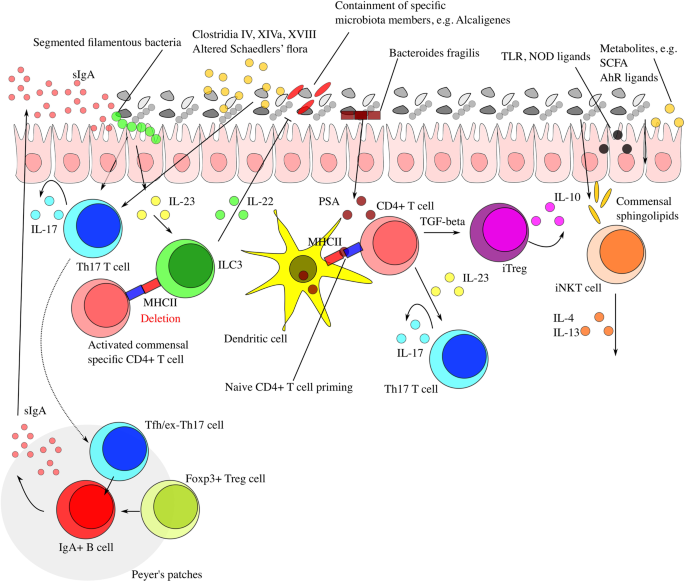
Interaction between microbiota and immunity in health and disease
Sender, R., Fuchs, S. & Milo, R. Are we really vastly outnumbered? revisiting the ratio of bacterial to host cells in humans. Cell 164, 337–340 (2016).
Google Scholar
Integrative HMP (iHMP) Research Network Consortium. The integrative human microbiome project. Nature 569, 641–648 (2019).
Google Scholar
Hacquard, S. et al. Microbiota and host nutrition across plant and animal kingdoms. Cell Host Microbe 17, 603–616 (2015).
Google Scholar
Lynch, J. B. & Hsiao, E. Y. Microbiomes as sources of emergent host phenotypes. Science 365, 1405–1409 (2019).
Google Scholar
Dethlefsen, L., McFall-Ngai, M. & Relman, D. A. An ecological and evolutionary perspective on human-microbe mutualism and disease. Nature 449, 811–818 (2007).
Google Scholar
Macpherson, A. J., Geuking, M. B. & McCoy, K. D. Immune responses that adapt the intestinal mucosa to commensal intestinal bacteria. Immunology 115, 153–162 (2005).
Google Scholar
Chu, H. & Mazmanian, S. K. Innate immune recognition of the microbiota promotes host-microbial symbiosis. Nat. Immunol. 14, 668–675 (2013).
Google Scholar
Zhang, M. et al. Interactions between intestinal microbiota and host immune response in inflammatory bowel disease. Front. Immunol. 8, 942 (2017).
Google Scholar
Valitutti, F., Cucchiara, S. & Fasano, A. Celiac disease and the microbiome. Nutrients 11, 2403 (2019).
Google Scholar
Maeda, Y. & Takeda, K. Host-microbiota interactions in rheumatoid arthritis. Exp. Mol. Med. 51, 150 (2019).
Google Scholar
Belizario, J. E., Faintuch, J. & Garay-Malpartida, M. Gut microbiome dysbiosis and immunometabolism: New frontiers for treatment of metabolic diseases. Mediators Inflamm. 2018, 2037838 (2018).
Google Scholar
Main, B. S. & Minter, M. R. Microbial immuno-communication in neurodegenerative diseases. Front. Neurosci. 11, 151 (2017).
Google Scholar
Gopalakrishnan, V., Helmink, B. A., Spencer, C. N., Reuben, A. & Wargo, J. A. The influence of the gut microbiome on cancer, immunity, and cancer immunotherapy. Cancer Cell 33, 570–580 (2018).
Google Scholar
Maynard, C. L., Elson, C. O., Hatton, R. D. & Weaver, C. T. Reciprocal interactions of the intestinal microbiota and immune system. Nature 489, 231–241 (2012).
Google Scholar
Belkaid, Y. & Harrison, O. J. Homeostatic immunity and the microbiota. Immunity 46, 562–576 (2017).
Google Scholar
Belkaid, Y. & Hand, T. W. Role of the microbiota in immunity and inflammation. Cell 157, 121–141 (2014).
Google Scholar
Gensollen, T., Iyer, S. S., Kasper, D. L. & Blumberg, R. S. How colonization by microbiota in early life shapes the immune system. Science 352, 539–544 (2016).
Google Scholar
Backhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 690–703 (2015).
Google Scholar
Koenig, J. E. et al. Succession of microbial consortia in the developing infant gut microbiome. Proc. Natl. Acad. Sci. USA 108(Suppl 1), 4578–4585 (2011).
Google Scholar
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486, 222–227 (2012).
Google Scholar
Russell, S. L. et al. Early life antibiotic-driven changes in microbiota enhance susceptibility to allergic asthma. EMBO Rep. 13, 440–447 (2012).
Google Scholar
Zhang, X., Zhivaki, D. & Lo-Man, R. Unique aspects of the perinatal immune system. Nat. Rev. Immunol. 17, 495–507 (2017).
Google Scholar
Bhutta, Z. A. & Black, R. E. Global maternal, newborn, and child health – So near and yet so far. N. Engl. J. Med. 369, 2226–2235 (2013).
Google Scholar
Neu, J. & Walker, W. A. Necrotizing enterocolitis. N. Engl. J. Med. 364, 255–264 (2011).
Google Scholar
Wang, J. et al. Dysbiosis of maternal and neonatal microbiota associated with gestational diabetes mellitus. Gut 67, 1614–1625 (2018).
Google Scholar
Gomez de Aguero, M. et al. The maternal microbiota drives early postnatal innate immune development. Science 351, 1296–1302 (2016).
Google Scholar
Dominguez-Bello, M. G. et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc. Natl. Acad. Sci. USA 107, 11971–11975 (2010).
Google Scholar
Caballero-Flores, G. et al. Maternal immunization confers protection to the offspring against an attaching and effacing pathogen through delivery of IgG in breast milk. Cell Host Microbe 25, 313–323 (2019).
Google Scholar
Zheng, W. et al. Microbiota-targeted maternal antibodies protect neonates from enteric infection. Nature 577, 543–548 (2020).
Google Scholar
Bauer, H., Horowitz, R. E., Levenson, S. M. & Popper, H. The response of the lymphatic tissue to the microbial flora. Studies on germfree mice. Am. J. Pathol. 42, 471–483 (1963).
Google Scholar
Umesaki, Y., Setoyama, H., Matsumoto, S. & Okada, Y. Expansion of alpha beta T-cell receptor-bearing intestinal intraepithelial lymphocytes after microbial colonization in germ-free mice and its independence from thymus. Immunology 79, 32–37 (1993).
Google Scholar
Hapfelmeier, S. et al. Reversible microbial colonization of germ-free mice reveals the dynamics of IgA immune responses. Science 328, 1705–1709 (2010).
Google Scholar
Ivanov, I. I. et al. Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe 4, 337–349 (2008).
Google Scholar
Ivanov, I. I. et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139, 485–498 (2009).
Google Scholar
Tan, T. G. et al. Identifying species of symbiont bacteria from the human gut that, alone, can induce intestinal Th17 cells in mice. Proc. Natl. Acad. Sci. USA 113, E8141–E8150 (2016).
Google Scholar
Atarashi, K. et al. Th17 cell induction by adhesion of microbes to intestinal epithelial cells. Cell 163, 367–380 (2015).
Google Scholar
Mazmanian, S. K., Liu, C. H., Tzianabos, A. O. & Kasper, D. L. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell 122, 107–118 (2005).
Google Scholar
Wesemann, D. R. et al. Microbial colonization influences early B-lineage development in the gut lamina propria. Nature 501, 112–115 (2013).
Google Scholar
Cahenzli, J., Koller, Y., Wyss, M., Geuking, M. B. & McCoy, K. D. Intestinal microbial diversity during early-life colonization shapes long-term IgE levels. Cell Host Microbe 14, 559–570 (2013).
Google Scholar
Fulde, M. et al. Neonatal selection by Toll-like receptor 5 influences long-term gut microbiota composition. Nature 560, 489–493 (2018).
Google Scholar
Mowat, A. M. To respond or not to respond – a personal perspective of intestinal tolerance. Nat. Rev. Immunol. 18, 405–415 (2018).
Google Scholar
Konrad, A., Cong, Y., Duck, W., Borlaza, R. & Elson, C. O. Tight mucosal compartmentation of the murine immune response to antigens of the enteric microbiota. Gastroenterology 130, 2050–2059 (2006).
Google Scholar
Belkaid, Y. & Naik, S. Compartmentalized and systemic control of tissue immunity by commensals. Nat. Immunol. 14, 646–653 (2013).
Google Scholar
Shan, M. et al. Mucus enhances gut homeostasis and oral tolerance by delivering immunoregulatory signals. Science 342, 447–453 (2013).
Google Scholar
Bansal, T., Alaniz, R. C., Wood, T. K. & Jayaraman, A. The bacterial signal indole increases epithelial-cell tight-junction resistance and attenuates indicators of inflammation. Proc. Natl. Acad. Sci. USA 107, 228–233 (2010).
Google Scholar
Peterson, D. A., McNulty, N. P., Guruge, J. L. & Gordon, J. I. IgA response to symbiotic bacteria as a mediator of gut homeostasis. Cell Host Microbe 2, 328–339 (2007).
Google Scholar
Macpherson, A. J. & Uhr, T. Induction of protective IgA by intestinal dendritic cells carrying commensal bacteria. Science 303, 1662–1665 (2004).
Google Scholar
Bevins, C. L. & Salzman, N. H. Paneth cells, antimicrobial peptides and maintenance of intestinal homeostasis. Nat. Rev. Microbiol. 9, 356–368 (2011).
Google Scholar
Ehmann, D. et al. Paneth cell α-defensins HD-5 and HD-6 display differential degradation into active antimicrobial fragments. Proc. Natl. Acad. Sci. USA 116, 3746–3751 (2019).
Google Scholar
Ahuja, M. et al. Orai1-mediated antimicrobial secretion from pancreatic acini shapes the gut microbiome and regulates gut innate immunity. Cell Metab. 25, 635–646 (2017).
Google Scholar
Rakoff-Nahoum, S., Paglino, J., Eslami-Varzaneh, F., Edberg, S. & Medzhitov, R. Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis. Cell 118, 229–241 (2004).
Google Scholar
Price, A. E. et al. A map of Toll-like receptor expression in the intestinal epithelium reveals distinct spatial, cell type-specific, and temporal patterns. Immunity 49, 560–575 (2018).
Google Scholar
Carvalho, F. A. et al. Transient inability to manage proteobacteria promotes chronic gut inflammation in TLR5-deficient mice. Cell Host Microbe 12, 139–152 (2012).
Google Scholar
Vijay-Kumar, M. et al. Metabolic syndrome and altered gut microbiota in mice lacking Toll-like receptor 5. Science 328, 228–231 (2010).
Google Scholar
Ubeda, C. et al. Familial transmission rather than defective innate immunity shapes the distinct intestinal microbiota of TLR-deficient mice. J. Exp. Med. 209, 1445–1456 (2012).
Google Scholar
Wen, L. et al. Innate immunity and intestinal microbiota in the development of type 1 diabetes. Nature 455, 1109–1113 (2008).
Google Scholar
Mazmanian, S. K., Round, J. L. & Kasper, D. L. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 453, 620–625 (2008).
Google Scholar
Lee, Y. K. et al. The protective role of Bacteroides fragilis in a murine model of colitis-associated colorectal cancer. mSphere 3, e00587–18 (2018).
Google Scholar
Ramakrishna, C. et al. Bacteroides fragilis polysaccharide A induces IL-10 secreting B and T cells that prevent viral encephalitis. Nat. Commun. 10, 2153 (2019).
Google Scholar
Erturk-Hasdemir, D. et al. Symbionts exploit complex signaling to educate the immune system. Proc. Natl. Acad. Sci. USA (2019).
Google Scholar
Brown, G. D. Dectin-1: a signalling non-TLR pattern-recognition receptor. Nat. Rev. Immunol. 6, 33–43 (2006).
Google Scholar
Tang, C. et al. Inhibition of Dectin-1 signaling ameliorates colitis by inducing Lactobacillus-mediated regulatory T cell expansion in the intestine. Cell Host Microbe 18, 183–197 (2015).
Google Scholar
Bouskra, D. et al. Lymphoid tissue genesis induced by commensals through NOD1 regulates intestinal homeostasis. Nature 456, 507–510 (2008).
Google Scholar
Ramanan, D., Tang, M. S., Bowcutt, R., Loke, P. & Cadwell, K. Bacterial sensor Nod2 prevents inflammation of the small intestine by restricting the expansion of the commensal Bacteroides vulgatus. Immunity 41, 311–324 (2014).
Google Scholar
Nigro, G., Rossi, R., Commere, P. H., Jay, P. & Sansonetti, P. J. The cytosolic bacterial peptidoglycan sensor Nod2 affords stem cell protection and links microbes to gut epithelial regeneration. Cell Host Microbe 15, 792–798 (2014).
Google Scholar
Janeway, C. A. Jr. & Medzhitov, R. Innate immune recognition. Annu. Rev. Immunol. 20, 197–216 (2002).
Google Scholar
Vaishnava, S. et al. The antibacterial lectin RegIIIgamma promotes the spatial segregation of microbiota and host in the intestine. Science 334, 255–258 (2011).
Google Scholar
Wang, S. et al. MyD88 adaptor-dependent microbial sensing by regulatory T cells promotes mucosal tolerance and enforces commensalism. Immunity 43, 289–303 (2015).
Google Scholar
Broz, P. & Dixit, V. M. Inflammasomes: Mechanism of assembly, regulation and signalling. Nat. Rev. Immunol. 16, 407–420 (2016).
Google Scholar
Elinav, E. et al. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 145, 745–757 (2011).
Google Scholar
Levy, M. et al. Microbiota-modulated metabolites shape the intestinal microenvironment by regulating NLRP6 inflammasome signaling. Cell 163, 1428–1443 (2015).
Google Scholar
Wlodarska, M. et al. NLRP6 inflammasome orchestrates the colonic host-microbial interface by regulating goblet cell mucus secretion. Cell 156, 1045–1059 (2014).
Google Scholar
Birchenough, G. M., Nystrom, E. E., Johansson, M. E. & Hansson, G. C. A sentinel goblet cell guards the colonic crypt by triggering Nlrp6-dependent Muc2 secretion. Science 352, 1535–1542 (2016).
Google Scholar
Wang, P. et al. Nlrp6 regulates intestinal antiviral innate immunity. Science 350, 826–830 (2015).
Google Scholar
Gálvez, E. J. C., Iljazovic, A., Gronow, A., Flavell, R. & Strowig, T. Shaping of intestinal microbiota in Nlrp6- and Rag2-deficient mice depends on community structure. Cell Rep. 21, 3914–3926 (2017).
Google Scholar
Castro-Dopico, T. et al. Anti-commensal IgG drives intestinal inflammation and type 17 immunity in ulcerative colitis. Immunity 50, 1099–1114 (2019).
Google Scholar
Seo, S. U. et al. Distinct commensals induce interleukin-1beta via NLRP3 inflammasome in inflammatory monocytes to promote intestinal inflammation in response to injury. Immunity 42, 744–755 (2015).
Google Scholar
Wolf, A. J. & Underhill, D. M. Peptidoglycan recognition by the innate immune system. Nat. Rev. Immunol. 18, 243–254 (2018).
Google Scholar
Ratsimandresy, R. A., Indramohan, M., Dorfleutner, A. & Stehlik, C. The AIM2 inflammasome is a central regulator of intestinal homeostasis through the IL-18/IL-22/STAT3 pathway. Cell Mol. Immunol. 14, 127–142 (2017).
Google Scholar
Saha, S. et al. Peptidoglycan recognition proteins protect mice from experimental colitis by promoting normal gut flora and preventing induction of interferon-gamma. Cell Host Microbe 8, 147–162 (2010).
Google Scholar
Jing, X. et al. Peptidoglycan recognition protein 3 and Nod2 synergistically protect mice from dextran sodium sulfate-induced colitis. J. Immunol. 193, 3055–3069 (2014).
Google Scholar
Franchi, L. et al. Cytosolic flagellin requires Ipaf for activation of caspase-1 and interleukin 1beta in salmonella-infected macrophages. Nat. Immunol. 7, 576–582 (2006).
Google Scholar
Zhu, H. et al. RNA virus receptor Rig-I monitors gut microbiota and inhibits colitis-associated colorectal cancer. J. Exp. Clin. Cancer Res. 36, 2 (2017).
Google Scholar
Hornung, V., Hartmann, R., Ablasser, A. & Hopfner, K. P. OAS proteins and cGAS: unifying concepts in sensing and responding to cytosolic nucleic acids. Nat. Rev. Immunol. 14, 521–528 (2014).
Google Scholar
Chudnovskiy, A. et al. Host-protozoan interactions protect from mucosal infections through activation of the inflammasome. Cell 167, 444–456 (2016).
Google Scholar
Mosser, D. M. & Edwards, J. P. Exploring the full spectrum of macrophage activation. Nat. Rev. Immunol. 8, 958–969 (2008).
Google Scholar
Danne, C. et al. A large polysaccharide produced by Helicobacter hepaticus induces an anti-inflammatory gene signature in macrophages. Cell Host Microbe 22, 733–745 (2017).
Google Scholar
Schulthess, J. et al. The short chain fatty acid butyrate imprints an antimicrobial program in macrophages. Immunity 50, 432–445 (2019).
Google Scholar
Wu, K. et al. Gut microbial metabolite trimethylamine N-oxide aggravates GVHD by inducing M1 macrophage polarization in mice. Blood (2020).
Google Scholar
Constantinides, M. G., McDonald, B. D., Verhoef, P. A. & Bendelac, A. A committed precursor to innate lymphoid cells. Nature 508, 397–401 (2014).
Google Scholar
Gury-BenAri, M. et al. The spectrum and regulatory landscape of intestinal innate lymphoid cells are shaped by the microbiome. Cell 166, 1231–1246 (2016).
Google Scholar
Sonnenberg, G. F. & Hepworth, M. R. Functional interactions between innate lymphoid cells and adaptive immunity. Nat. Rev. Immunol. 19, 599–613 (2019).
Google Scholar
McDonald, B. D., Jabri, B. & Bendelac, A. Diverse developmental pathways of intestinal intraepithelial lymphocytes. Nat. Rev. Immunol. 18, 514–525 (2018).
Google Scholar
Chun, E. et al. Metabolite-sensing receptor Ffar2 regulates colonic group 3 innate lymphoid cells and gut immunity. Immunity 51, 871–884 (2019).
Google Scholar
Bostick, J. W. et al. Dichotomous regulation of group 3 innate lymphoid cells by nongastric Helicobacter species. Proc. Natl. Acad. Sci. USA 116, 24760–24769 (2019).
Google Scholar
Guo, X. et al. Innate lymphoid cells control early colonization resistance against intestinal pathogens through ID2-dependent regulation of the microbiota. Immunity 42, 731–743 (2015).
Google Scholar
Rankin, L. C. et al. Complementarity and redundancy of IL-22-producing innate lymphoid cells. Nat. Immunol. 17, 179–186 (2016).
Google Scholar
Chua, H. H. et al. Intestinal dysbiosis featuring abundance of Ruminococcus gnavus associates with allergic diseases in infants. Gastroenterology 154, 154–167 (2018).
Google Scholar
Sterlin, D. et al. Human IgA binds a diverse array of commensal bacteria. J. Exp. Med. 217, e20181635 (2020).
Google Scholar
Sutherland, D. B., Suzuki, K. & Fagarasan, S. Fostering of advanced mutualism with gut microbiota by immunoglobulin A. Immunol. Rev. 270, 20–31 (2016).
Google Scholar
Kawamoto, S. et al. Foxp3+ T cells regulate immunoglobulin A selection and facilitate diversification of bacterial species responsible for immune homeostasis. Immunity 41, 152–165 (2014).
Google Scholar
Palm, N. W. et al. Immunoglobulin A coating identifies colitogenic bacteria in inflammatory bowel disease. Cell 158, 1000–1010 (2014).
Google Scholar
Shulzhenko, N. et al. Crosstalk between B lymphocytes, microbiota and the intestinal epithelium governs immunity versus metabolism in the gut. Nat. Med. 17, 1585–1593 (2011).
Google Scholar
Nagashima, K. et al. Identification of subepithelial mesenchymal cells that induce IgA and diversify gut microbiota. Nat. Immunol. 18, 675–682 (2017).
Google Scholar
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455 (2013).
Google Scholar
Atarashi, K. et al. Induction of colonic regulatory T cells by indigenous clostridium species. Science 331, 334–337 (2011).
Google Scholar
Smith, P. M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013).
Google Scholar
Hegazy, A. N. et al. Circulating and tissue-resident CD4+ T cells with reactivity to intestinal microbiota are abundant in healthy individuals and function is altered during inflammation. Gastroenterology 153, 1320–1337 (2017).
Google Scholar
Miossec, P. & Kolls, J. K. Targeting IL-17 and Th17 cells in chronic inflammation. Nat. Rev. Drug Discov. 11, 763–776 (2012).
Google Scholar
Omenetti, S. et al. The intestine harbors functionally distinct homeostatic tissue-resident and inflammatory Th17 cells. Immunity 51, 77–89 (2019).
Google Scholar
Naik, S. et al. Compartmentalized control of skin immunity by resident commensals. Science 337, 1115–1119 (2012).
Google Scholar
Dutzan, N. et al. On-going mechanical damage from mastication drives homeostatic Th17 cell responses at the oral barrier. Immunity 46, 133–147 (2017).
Google Scholar
Bedoui, S., Heath, W. R. & Mueller, S. N. CD4(+) T-cell help amplifies innate signals for primary CD8(+) T-cell immunity. Immunol. Rev. 272, 52–64 (2016).
Google Scholar
Bachem, A. et al. Microbiota-derived short-chain fatty acids promote the memory potential of antigen-activated CD8(+) T cells. Immunity 51, 285–297 (2019).
Google Scholar
Song, X. et al. Microbial bile acid metabolites modulate gut RORgamma(+) regulatory T cell homeostasis. Nature 577, 410–415 (2020).
Google Scholar
Crotty, S. T follicular helper cell differentiation, function, and roles in disease. Immunity 41, 529–542 (2014).
Google Scholar
Kawamoto, S. et al. The inhibitory receptor PD-1 regulates IgA selection and bacterial composition in the gut. Science 336, 485–489 (2012).
Google Scholar
Proietti, M. et al. ATP-gated ionotropic P2X7 receptor controls follicular T helper cell numbers in Peyer’s patches to promote host-microbiota mutualism. Immunity 41, 789–801 (2014).
Google Scholar
Kubinak, J. L. et al. MyD88 signaling in T cells directs IgA-mediated control of the microbiota to promote health. Cell Host Microbe 17, 153–163 (2015).
Google Scholar
Teng, F. et al. Gut microbiota drive autoimmune arthritis by promoting differentiation and migration of Peyer’s patch T follicular helper cells. Immunity 44, 875–888 (2016).
Google Scholar
Rescigno, M., Rotta, G., Valzasina, B. & Ricciardi-Castagnoli, P. Dendritic cells shuttle microbes across gut epithelial monolayers. Immunobiology 204, 572–581 (2001).
Google Scholar
Martinez-Lopez, M. et al. Microbiota sensing by Mincle-Syk axis in dendritic cells regulates interleukin-17 and -22 production and promotes intestinal barrier integrity. Immunity 50, 446–461 (2019).
Google Scholar
Jie, Z. et al. NIK signaling axis regulates dendritic cell function in intestinal immunity and homeostasis. Nat. Immunol. 19, 1224–1235 (2018).
Google Scholar
Wingender, G. et al. Neutrophilic granulocytes modulate invariant NKT cell function in mice and humans. J. Immunol. 188, 3000–3008 (2012).
Google Scholar
An, D. et al. Sphingolipids from a symbiotic microbe regulate homeostasis of host intestinal natural killer T cells. Cell 156, 123–133 (2014).
Google Scholar
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
Google Scholar
Vojdani, A. A potential link between environmental triggers and autoimmunity. Autoimmune Dis. 2014, 437231 (2014).
Google Scholar
Yamamoto-Hanada, K., Yang, L., Narita, M., Saito, H. & Ohya, Y. Influence of antibiotic use in early childhood on asthma and allergic diseases at age 5. Ann. Allergy Asthma Immunol. 119, 54–58 (2017).
Google Scholar
Becattini, S., Taur, Y. & Pamer, E. G. Antibiotic-induced changes in the intestinal microbiota and disease. Trends Mol. Med. 22, 458–478 (2016).
Google Scholar
Sato, H. et al. Antibiotics suppress activation of intestinal mucosal mast cells and reduce dietary lipid absorption in Sprague-Dawley rats. Gastroenterology 151, 923–932 (2016).
Google Scholar
Scott, N. A. et al. Antibiotics induce sustained dysregulation of intestinal T cell immunity by perturbing macrophage homeostasis. Sci. Transl. Med. 10, eaao4755 (2018).
Google Scholar
Kim, Y. G. et al. Gut dysbiosis promotes M2 macrophage polarization and allergic airway inflammation via fungi-induced PGE(2). Cell Host Microbe 15, 95–102 (2014).
Google Scholar
Kim, M. et al. Critical Role for the microbiota in CX3CR1(+) intestinal mononuclear phagocyte regulation of intestinal T cell responses. Immunity 49, 151–163 (2018).
Google Scholar
Ohnmacht, C. et al. MUCOSAL IMMUNOLOGY. The microbiota regulates type 2 immunity through RORgammat(+) T cells. Science 349, 989–993 (2015).
Google Scholar
Hagan, T. et al. Antibiotics-driven gut microbiome perturbation alters immunity to vaccines in humans. Cell 178, 1313–1328 (2019).
Google Scholar
Christ, A., Lauterbach, M. & Latz, E. Western diet and the immune system: An inflammatory connection. Immunity 51, 794–811 (2019).
Google Scholar
Devkota, S. et al. Dietary-fat-induced taurocholic acid promotes pathobiont expansion and colitis in Il10−/− mice. Nature 487, 104–108 (2012).
Google Scholar
Cheng, L. et al. High fat diet exacerbates dextran sulfate sodium induced colitis through disturbing mucosal dendritic cell homeostasis. Int. Immunopharmacol. 40, 1–10 (2016).
Google Scholar
Haghikia, A. et al. Dietary fatty acids directly impact central nervous system autoimmunity via the small intestine. Immunity 43, 817–829 (2015).
Google Scholar
He, B. et al. Resetting microbiota by Lactobacillus reuteri inhibits T reg deficiency-induced autoimmunity via adenosine A2A receptors. J. Exp. Med. 214, 107–123 (2017).
Google Scholar
Rodriguez-Palacios, A. et al. The artificial sweetener splenda promotes gut Proteobacteria, dysbiosis, and myeloperoxidase reactivity in Crohn’s disease-like ileitis. Inflamm. Bowel Dis. 24, 1005–1020 (2018).
Google Scholar
Viennois, E., Merlin, D., Gewirtz, A. T. & Chassaing, B. Dietary emulsifier-induced low-grade inflammation promotes colon carcinogenesis. Cancer Res. 77, 27–40 (2017).
Google Scholar
Martinez, I. et al. Gut microbiome composition is linked to whole grain-induced immunological improvements. ISME J. 7, 269–280 (2013).
Google Scholar
Cignarella, F. et al. Intermittent fasting confers protection in CNS autoimmunity by altering the gut microbiota. Cell Metab. 27, 1222–1235 (2018).
Google Scholar
Rangan, P. et al. Fasting-mimicking diet modulates microbiota and promotes intestinal regeneration to reduce inflammatory bowel disease pathology. Cell Rep. 26, 2704–2719 (2019).
Google Scholar
Bishehsari, F. et al. Abnormal eating patterns cause circadian disruption and promote alcohol-associated colon carcinogenesis. Cell Mol. Gastroenterol. Hepatol. 9, 219–237 (2020).
Google Scholar
Rosshart, S. P. et al. Laboratory mice born to wild mice have natural microbiota and model human immune responses. Science 365, eaaw4361 (2019).
Google Scholar
Kaplan, G. G. The global burden of IBD: from 2015 to 2025. Nat. Rev. Gastroenterol. Hepatol. 12, 720–727 (2015).
Google Scholar
Kostic, A. D., Xavier, R. J. & Gevers, D. The microbiome in inflammatory bowel disease: current status and the future ahead. Gastroenterology 146, 1489–1499 (2014).
Google Scholar
Gevers, D. et al. The treatment-naive microbiome in new-onset Crohn’s disease. Cell Host Microbe 15, 382–392 (2014).
Google Scholar
Franzosa, E. A. et al. Gut microbiome structure and metabolic activity in inflammatory bowel disease. Nat. Microbiol. 4, 293–305 (2019).
Google Scholar
Lloyd-Price, J. et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature 569, 655–662 (2019).
Google Scholar
de Souza, H. S. & Fiocchi, C. Immunopathogenesis of IBD: current state of the art. Nat. Rev. Gastroenterol. Hepatol. 13, 13–27 (2016).
Google Scholar
Martini, E., Krug, S. M., Siegmund, B., Neurath, M. F. & Becker, C. Mend your fences: The epithelial barrier and its relationship with mucosal immunity in inflammatory bowel disease. Cell Mol. Gastroenterol. Hepatol. 4, 33–46 (2017).
Google Scholar
Van der Sluis, M. et al. Muc2-deficient mice spontaneously develop colitis, indicating that MUC2 is critical for colonic protection. Gastroenterology 131, 117–129 (2006).
Google Scholar
Liso, M. et al. A specific mutation in Muc2 determines early dysbiosis in colitis-prone Winnie mice. Inflamm. Bowel Dis. 26, 546–556 (2020).
Google Scholar
Ogura, Y. et al. A frameshift mutation in NOD2 associated with susceptibility to Crohn’s disease. Nature 411, 603–606 (2001).
Google Scholar
Hugot, J. P. et al. Association of NOD2 leucine-rich repeat variants with susceptibility to Crohn’s disease. Nature 411, 599–603 (2001).
Google Scholar
Petnicki-Ocwieja, T. et al. Nod2 is required for the regulation of commensal microbiota in the intestine. Proc. Natl. Acad. Sci. USA 106, 15813–15818 (2009).
Google Scholar
Cadwell, K. et al. A key role for autophagy and the autophagy gene Atg16l1 in mouse and human intestinal Paneth cells. Nature 456, 259–263 (2008).
Google Scholar
Aden, K. et al. ATG16L1 orchestrates interleukin-22 signaling in the intestinal epithelium via cGAS-STING. J. Exp. Med. 215, 2868–2886 (2018).
Google Scholar
Seregin, S. S. et al. NLRP6 protects Il10(−/−) mice from colitis by limiting colonization of Akkermansia muciniphila. Cell Rep. 19, 733–745 (2017).
Google Scholar
Schaubeck, M. et al. Dysbiotic gut microbiota causes transmissible Crohn’s disease-like ileitis independent of failure in antimicrobial defence. Gut 65, 225–237 (2016).
Google Scholar
Britton, G. J. et al. Microbiotas from humans with inflammatory bowel disease alter the balance of gut Th17 and RORgammat(+) regulatory T cells and exacerbate colitis in mice. Immunity 50, 212–224 (2019).
Google Scholar
Caruso, R. et al. A specific gene-microbe interaction drives the development of Crohn’s disease-like colitis in mice. Sci. Immunol. 4, eaaw4341 (2019).
Google Scholar
Atarashi, K. et al. Ectopic colonization of oral bacteria in the intestine drives TH1 cell induction and inflammation. Science 358, 359–365 (2017).
Google Scholar
Torres, J. et al. Infants born to mothers with IBD present with altered gut microbiome that transfers abnormalities of the adaptive immune system to germ-free mice. Gut 69, 42–51 (2020).
Google Scholar
Scher, J. U. et al. Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. Elife 2, e01202 (2013).
Google Scholar
Maeda, Y. et al. Dysbiosis contributes to arthritis development via activation of autoreactive T cells in the intestine. Arthritis Rheumatol. 68, 2646–2661 (2016).
Google Scholar
Alpizar-Rodriguez, D. et al. Prevotella copri in individuals at risk for rheumatoid arthritis. Ann. Rheum. Dis. 78, 590–593 (2019).
Google Scholar
Chen, J. et al. An expansion of rare lineage intestinal microbes characterizes rheumatoid arthritis. Genome Med. 8, 43 (2016).
Google Scholar
Zhang, X. et al. The oral and gut microbiomes are perturbed in rheumatoid arthritis and partly normalized after treatment. Nat. Med. 21, 895–905 (2015).
Google Scholar
Wang, Q. & Xu, R. Data-driven multiple-level analysis of gut-microbiome-immune-joint interactions in rheumatoid arthritis. BMC Genom. 20, 124 (2019).
Google Scholar
Abdollahi-Roodsaz, S. et al. Stimulation of TLR2 and TLR4 differentially skews the balance of T cells in a mouse model of arthritis. J. Clin. Invest. 118, 205–216 (2008).
Google Scholar
Rogier, R. et al. Aberrant intestinal microbiota due to IL-1 receptor antagonist deficiency promotes IL-17- and TLR4-dependent arthritis. Microbiome 5, 63 (2017).
Google Scholar
Wu, H. J. et al. Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity 32, 815–827 (2010).
Google Scholar
de Aquino, S. G. et al. Periodontal pathogens directly promote autoimmune experimental arthritis by inducing a TLR2- and IL-1-driven Th17 response. J. Immunol. 192, 4103–4111 (2014).
Google Scholar
Hotamisligil, G. S. Inflammation, metaflammation and immunometabolic disorders. Nature 542, 177–185 (2017).
Google Scholar
Tilg, H., Zmora, N., Adolph, T. E. & Elinav, E. The intestinal microbiota fuelling metabolic inflammation. Nat. Rev. Immunol. 20, 40–54 (2020).
Google Scholar
Kolodziejczyk, A. A., Zheng, D., Shibolet, O. & Elinav, E. The role of the microbiome in NAFLD and NASH. EMBO Mol. Med. 11, e9302 (2019).
Google Scholar
Henao-Mejia, J. et al. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature 482, 179–185 (2012).
Google Scholar
Bodogai, M. et al. Commensal bacteria contribute to insulin resistance in aging by activating innate B1a cells. Sci. Transl. Med. 10, eaat4271 (2018).
Google Scholar
Virtue, A. T. et al. The gut microbiota regulates white adipose tissue inflammation and obesity via a family of microRNAs. Sci. Transl. Med. 11, eaav1892 (2019).
Google Scholar
Truax, A. D. et al. The inhibitory innate immune sensor NLRP12 maintains a threshold against obesity by regulating gut microbiota homeostasis. Cell Host Microbe 24, 364–378 (2018).
Google Scholar
Koeth, R. A. et al. l-Carnitine in omnivorous diets induces an atherogenic gut microbial pathway in humans. J. Clin. Invest. 129, 373–387 (2019).
Google Scholar
Wang, Z. et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472, 57–63 (2011).
Google Scholar
Gur, C. et al. Binding of the Fap2 protein of Fusobacterium nucleatum to human inhibitory receptor TIGIT protects tumors from immune cell attack. Immunity 42, 344–355 (2015).
Google Scholar
Mima, K. et al. Fusobacterium nucleatum and T cells in colorectal carcinoma. JAMA Oncol. 1, 653–661 (2015).
Google Scholar
Ma, C. et al. Gut microbiome-mediated bile acid metabolism regulates liver cancer via NKT cells. Science 360, eaan5931 (2018).
Google Scholar
Matson, V. et al. The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science 359, 104–108 (2018).
Google Scholar
Gopalakrishnan, V. et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359, 97–103 (2018).
Google Scholar
Sivan, A. et al. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science 350, 1084–1089 (2015).
Google Scholar
Routy, B. et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 359, 91–97 (2018).
Google Scholar
Vetizou, M. et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079–1084 (2015).
Google Scholar
Viaud, S. et al. The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science 342, 971–976 (2013).
Google Scholar
Zitvogel, L., Ayyoub, M., Routy, B. & Kroemer, G. Microbiome and anticancer immunosurveillance. Cell 165, 276–287 (2016).
Google Scholar
Pushalkar, S. et al. The pancreatic cancer microbiome promotes oncogenesis by induction of innate and adaptive immune suppression. Cancer Discov. 8, 403–416 (2018).
Google Scholar
Geller, L. T. et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science 357, 1156–1160 (2017).
Google Scholar
Riquelme, E. et al. Tumor microbiome diversity and composition influence pancreatic cancer outcomes. Cell 178, 795–806 (2019).
Google Scholar
Grice, E. A. et al. Topographical and temporal diversity of the human skin microbiome. Science 324, 1190–1192 (2009).
Google Scholar
Oh, J., Byrd, A. L., Park, M., Kong, H. H. & Segre, J. A. Temporal stability of the human skin microbiome. Cell 165, 854–866 (2016).
Google Scholar
Oh, J. et al. Biogeography and individuality shape function in the human skin metagenome. Nature 514, 59–64 (2014).
Google Scholar
Chehoud, C. et al. Complement modulates the cutaneous microbiome and inflammatory milieu. Proc. Natl. Acad. Sci. USA 110, 15061–15066 (2013).
Google Scholar
Brandwein, M., Bentwich, Z. & Steinberg, D. Endogenous antimicrobial peptide expression in response to bacterial epidermal colonization. Front. Immunol. 8, 1637 (2017).
Google Scholar
Naik, S. et al. Commensal-dendritic-cell interaction specifies a unique protective skin immune signature. Nature 520, 104–108 (2015).
Google Scholar
Linehan, J. L. et al. Non-classical immunity controls microbiota impact on skin immunity and tissue repair. Cell 172, 784–796 (2018).
Google Scholar
Lai, Y. et al. Commensal bacteria regulate Toll-like receptor 3-dependent inflammation after skin injury. Nat. Med. 15, 1377–1382 (2009).
Google Scholar
Scharschmidt, T. C. et al. A wave of regulatory T cells into neonatal skin mediates tolerance to commensal microbes. Immunity 43, 1011–1021 (2015).
Google Scholar
Scharschmidt, T. C. et al. Commensal microbes and hair follicle morphogenesis coordinately drive Treg migration into neonatal skin. Cell Host Microbe 21, 467–477 (2017).
Google Scholar
Sanford, J. A. et al. Inhibition of HDAC8 and HDAC9 by microbial short-chain fatty acids breaks immune tolerance of the epidermis to TLR ligands. Sci. Immunol. 1, eaah4609 (2016).
Google Scholar
Nakatsuji, T. et al. Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Sci. Transl. Med. 9, eaah4680 (2017).
Google Scholar
Kong, H. H. et al. Temporal shifts in the skin microbiome associated with disease flares and treatment in children with atopic dermatitis. Genome Res. 22, 850–859 (2012).
Google Scholar
Stehlikova, Z. et al. Dysbiosis of skin microbiota in Psoriatic patients: co-occurrence of fungal and bacterial communities. Front. Microbiol. 10, 438 (2019).
Google Scholar
Belkaid, Y. & Segre, J. A. Dialogue between skin microbiota and immunity. Science 346, 954–959 (2014).
Google Scholar
Nakamura, Y. et al. Staphylococcus delta-toxin induces allergic skin disease by activating mast cells. Nature 503, 397–401 (2013).
Google Scholar
Uluckan, O. et al. Cutaneous immune cell-microbiota interactions are controlled by epidermal JunB/AP-1. Cell Rep. 29, 844–859 (2019).
Google Scholar
Ichinohe, T. et al. Microbiota regulates immune defense against respiratory tract influenza A virus infection. Proc. Natl. Acad. Sci. USA 108, 5354–5359 (2011).
Google Scholar
Fagundes, C. T. et al. Transient TLR activation restores inflammatory response and ability to control pulmonary bacterial infection in germfree mice. J. Immunol. 188, 1411–1420 (2012).
Google Scholar
Trompette, A. et al. Dietary fiber confers protection against flu by shaping Ly6c(−) patrolling monocyte hematopoiesis and CD8(+) T cell metabolism. Immunity 48, 992–1005 (2018).
Google Scholar
Steed, A. L. et al. The microbial metabolite desaminotyrosine protects from influenza through type I interferon. Science 357, 498–502 (2017).
Google Scholar
Marsland, B. J. & Gollwitzer, E. S. Host-microorganism interactions in lung diseases. Nat. Rev. Immunol. 14, 827–835 (2014).
Google Scholar
Gollwitzer, E. S. et al. Lung microbiota promotes tolerance to allergens in neonates via PD-L1. Nat. Med. 20, 642–647 (2014).
Google Scholar
Pattaroni, C. et al. Early-life formation of the microbial and immunological environment of the human airways. Cell Host Microbe 24, 857–865 (2018).
Google Scholar
Dickson, R. P., Martinez, F. J. & Huffnagle, G. B. The role of the microbiome in exacerbations of chronic lung diseases. Lancet 384, 691–702 (2014).
Google Scholar
Yadava, K. et al. Microbiota promotes chronic pulmonary inflammation by enhancing IL-17A and autoantibodies. Am. J. Respir. Crit. Care Med. 193, 975–987 (2016).
Google Scholar
Segal, L. N. et al. Enrichment of the lung microbiome with oral taxa is associated with lung inflammation of a Th17 phenotype. Nat. Microbiol. 1, 16031 (2016).
Google Scholar
Larsen, J. M. et al. Chronic obstructive pulmonary disease and asthma-associated Proteobacteria, but not commensal Prevotella spp., promote Toll-like receptor 2-independent lung inflammation and pathology. Immunology 144, 333–342 (2015).
Google Scholar
Jin, C. et al. Commensal microbiota promote lung cancer development via gammadelta T cells. Cell 176, 998–1013 (2019).
Google Scholar
Corbitt, N. et al. Gut bacteria drive Kupffer cell expansion via MAMP-mediated ICAM-1 induction on sinusoidal endothelium and influence preservation-reperfusion injury after orthotopic liver transplantation. Am. J. Pathol. 182, 180–191 (2013).
Google Scholar
Chen, J. et al. Natural killer T cells play a necessary role in modulating of immune-mediated liver injury by gut microbiota. Sci. Rep. 4, 7259 (2014).
Google Scholar
Liang, S., Webb, T. & Li, Z. Probiotic antigens stimulate hepatic natural killer T cells. Immunology 141, 203–210 (2014).
Google Scholar
Paik, Y. H. et al. Toll-like receptor 4 mediates inflammatory signaling by bacterial lipopolysaccharide in human hepatic stellate cells. Hepatology 37, 1043–1055 (2003).
Google Scholar
Nakamoto, N. et al. Gut pathobionts underlie intestinal barrier dysfunction and liver T helper 17 cell immune response in primary sclerosing cholangitis. Nat. Microbiol. 4, 492–503 (2019).
Google Scholar
Liwinski, T. et al. Alterations of the bile microbiome in primary sclerosing cholangitis. Gut 69, 665–672 (2020).
Google Scholar
Yoshimoto, S. et al. Obesity-induced gut microbial metabolite promotes liver cancer through senescence secretome. Nature 499, 97–101 (2013).
Google Scholar
Loo, T. M. et al. Gut microbiota promotes obesity-associated liver cancer through PGE2-mediated suppression of antitumor immunity. Cancer Discov. 7, 522–538 (2017).
Google Scholar
Dapito, D. H. et al. Promotion of hepatocellular carcinoma by the intestinal microbiota and TLR4. Cancer Cell 21, 504–516 (2012).
Google Scholar
Sharon, G., Sampson, T. R., Geschwind, D. H. & Mazmanian, S. K. The central nervous system and the gut microbiome. Cell 167, 915–932 (2016).
Google Scholar
Butovsky, O. & Weiner, H. L. Microglial signatures and their role in health and disease. Nat. Rev. Neurosci. 19, 622–635 (2018).
Google Scholar
Erny, D. et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat. Neurosci. 18, 965–977 (2015).
Google Scholar
Matcovitch-Natan, O. et al. Microglia development follows a stepwise program to regulate brain homeostasis. Science 353, aad8670 (2016).
Google Scholar
Thion, M. S. et al. Microbiome influences prenatal and adult microglia in a sex-specific manner. Cell 172, 500–516 (2018).
Google Scholar
Abdel-Haq, R., Schlachetzki, J. C. M., Glass, C. K. & Mazmanian, S. K. Microbiome-microglia connections via the gut-brain axis. J. Exp. Med. 216, 41–59 (2019).
Google Scholar
Benakis, C. et al. Commensal microbiota affects ischemic stroke outcome by regulating intestinal gammadelta T cells. Nat. Med. 22, 516–523 (2016).
Google Scholar
Ochoa-Reparaz, J. et al. Role of gut commensal microflora in the development of experimental autoimmune encephalomyelitis. J. Immunol. 183, 6041–6050 (2009).
Google Scholar
Kim, S. et al. Maternal gut bacteria promote neurodevelopmental abnormalities in mouse offspring. Nature 549, 528–532 (2017).
Google Scholar
Reed, M. D. et al. IL-17a promotes sociability in mouse models of neurodevelopmental disorders. Nature 577, 249–253 (2020).
Google Scholar
Yang, D. et al. Dysregulated lung commensal bacteria drive interleukin-17B production to promote pulmonary fibrosis through their outer membrane vesicles. Immunity 50, 692–706 (2019).
Google Scholar
O’Callaghan, J. L. et al. Re-assessing microbiomes in the low-biomass reproductive niche. BJOG 127, 147–158 (2020).
Google Scholar
Minich, J. J. et al. KatharoSeq enables high-throughput microbiome analysis from low-biomass samples. mSystems 3, e00218–17 (2018).
Google Scholar
Salter, S. J. et al. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 12, 87 (2014).
Google Scholar
de Goffau, M. C. et al. Human placenta has no microbiome but can contain potential pathogens. Nature 572, 329–334 (2019).
Google Scholar
Kuperman, A. A. et al. Deep microbial analysis of multiple placentas shows no evidence for a placental microbiome. BJOG 127, 159–169 (2020).
Google Scholar
Karstens, L. et al. Controlling for contaminants in low-biomass 16S rRNA gene sequencing experiments. mSystems 4, e00290–19 (2019).
Google Scholar
Burnham, P. et al. Separating the signal from the noise in metagenomic cell-free DNA sequencing. Microbiome 8, 18 (2020).
Google Scholar
Eisenhofer, R. et al. Contamination in low microbial biomass microbiome studies: Issues and recommendations. Trends Microbiol. 27, 105–117 (2019).
Google Scholar
Limon, J. J. et al. Malassezia is associated with Crohn’s disease and exacerbates colitis in mouse models. Cell Host Microbe 25, 377–388 (2019).
Google Scholar
Sokol, H. et al. Fungal microbiota dysbiosis in IBD. Gut 66, 1039–1048 (2017).
Google Scholar
Norman, J. M. et al. Disease-specific alterations in the enteric virome in inflammatory bowel disease. Cell 160, 447–460 (2015).
Google Scholar
Zuo, T. et al. Gut mucosal virome alterations in ulcerative colitis. Gut 68, 1169–1179 (2019).
Google Scholar
Liu, T. C. & Stappenbeck, T. S. Genetics and pathogenesis of inflammatory bowel disease. Annu. Rev. Pathol. 11, 127–148 (2016).
Google Scholar
Rosshart, S. P. et al. Wild mouse gut microbiota promotes host fitness and improves disease resistance. Cell 171, 1015–1028 (2017).
Google Scholar
Tanoue, T. et al. A defined commensal consortium elicits CD8 T cells and anti-cancer immunity. Nature 565, 600–605 (2019).
Google Scholar
Zhu, W. et al. Precision editing of the gut microbiota ameliorates colitis. Nature 553, 208–211 (2018).
Google Scholar
Van Belleghem, J. D., Dabrowska, K., Vaneechoutte, M., Barr, J. J. & Bollyky, P. L. Interactions between bacteriophage, bacteria, and the mammalian immune system. Viruses 11, E10 (2018).
Google Scholar
Zeevi, D. et al. Personalized nutrition by prediction of glycemic responses. Cell 163, 1079–1094 (2015).
Google Scholar
Levy, M., Thaiss, C. A. & Elinav, E. Metabolites: messengers between the microbiota and the immune system. Genes Dev. 30, 1589–1597 (2016).
Google Scholar
Chen, H. et al. A forward chemical genetic screen reveals gut microbiota metabolites that modulate host physiology. Cell 177, 1217–1231 (2019).
Google Scholar
link