Materials and reagents
2-Hydroxyethyl methacrylate (HEMA), 3,4-dihydro-2H-pyran, octadecyl acrylate (ODA), benzoyl peroxide (BPO), 3, 3′, 5, 5’-tetramethylbenzidine (TMB), ammonium persulfate (APS), N,N,N′,N′-tetramethyl ethylenediamine (TEMED), and N,N′-methylenebis(acrylamide) (MBAA) were supplied by Aladdin Reagent Co., Ltd. o-Phenylenediamine (OPD) and tasquinimod (TAS) were purchased from Macklin Reagent Co., Ltd. p-Toluenesulfonic acid (PTSA) was obtained from Rhawn Reagent Co., Ltd. Phosphate buffer solution (PBS) was purchased from Sangon Biotech Co., Ltd. The silicone elastomer Sylgard 184 was obtained from Dow Corning (catalog no. 3097358-1004). Fibrinogen, Ethylene glycol dimethacrylate (EGDMA), streptozocin (STZ), and 2-methacryloyloxyethyl phosphorylcholine (MPC) were purchased from Sigma-Aldrich. TPU (Lubrizol Pellethane 2363-80a), POE (ENGAGE 8999), PA (Pebax MED 6333), and EMA (LOTADER ax8900) were purchased from corresponding companies and processed into 1 mm thick sheets. The Silhouette infusion set (Silhouette Paradigm MMT-381A, MiniMed Reservoir (no. MMT-332A) and Medtronic MiniMed 722 pump were purchased from GEMCO Medical.
Synthesis of HPEMA monomers
2-((tetrahydro-2H-pyran-2-yl)oxy)ethyl methacrylate (HPEMA) was synthesized as follows. To a stirred solution of HEMA (10 g, 1.0 eq.) in 30 mL of dry ether was added PTSA (1.3 g, 0.1 eq.). To this reaction mixture, a solution of 2,3-dihydropyran (10 mL, 1.4 eq.) in 5 mL dry ether was added dropwise at 0 °C. The reaction contents were warmed to room temperature and stirred in the dark for 3 h, concentrated and purified by column chromatography with ethyl acetate/hexane (1:4 v/v) to give the desired monomer as light-yellow oil in 87% yield (17.3 g). 1H and 13C NMR spectra were collected on a Bruker AVANCE III 500 spectrometer at 500 MHz and 125 MHz, respectively, using CDCl3 as the solvent. 1H NMR chemical shifts were referenced to the resonance for residual protonated solvent (δ 0.00 for TMS in CDCl3). 13C NMR chemical shifts were referenced to the solvent (δ 77.16 for CDCl3). 1H NMR (500 MHz, CDCl3) δ 6.07 (s, 1H), 5.52 (s, 1H), 4.60 (t, J = 3.4 Hz, 1H), 4.39–4.26 (m, 2H), 3.95 (ddd, J = 11.3, 6.1, 3.7 Hz, 1H), 3.91–3.80 (m, 1H), 3.69 (ddd, J = 11.3, 6.4, 3.7 Hz, 1H), 3.57–3.45 (m, 1H), 1.96 (s, 3H), 1.89–1.77 (m, 1H), 1.76–1.66 (m, 1H), 1.57 (dddd, J = 14.0, 11.8, 10.1, 5.2, 4H). 13C NMR (125 MHz, CDCl3) δ 167.38 (s), 136.24 (s), 125.64 (s), 98.67 (s), 65.10 (s), 63.89 (s), 61.94 (s), 30.45 (s), 25.41 (s), 19.19 (s), 18.32 (s). Mass spectra were acquired using an Agilent 6545 Q-TOF mass spectrometer. ESI-MS: [M+Na]+: 237.11.
Fabrication and characterization of EVADE materials
The EVADE materials were fabricated by radical bulk copolymerization in Poly tetra fluoroethylene (PTFE) molds with the desired shape. A reaction mixture containing HPEMA monomer, ODA monomer, and 0.5 wt% (relative to the total weight of monomers) of initiator BPO was thermo-polymerized using PTFE slides separated by a 1 mm thick PTFE spacer. The reaction was kept at 70 °C for 2 d. After polymerization, the elastomer samples were soaked thoroughly until equilibrium in anhydrous ethanol to remove residual monomer, followed by drying under vacuum at room temperature for further tests. H90 elastomers crosslinked with EGDMA were synthesized using a similar procedure. The amount of EGDMA added is from 0 to 1 wt%. The chemical structures of these EVADE materials (H0 to H100, 10% as a step) were confirmed by 1H NMR (400 MHz, CDCl3). The molecular structures of the EVADE materials were analyzed by Fourier Transform Infrared Spectroscopy (FTIR) using a Thermo Nicolet 6700 FT-IR Spectrometer with a Smart iTR Attenuated Total Reflectance (ATR). The wavenumber ranged from 500 to 4000 cm−1 with 32 scans.
XRD
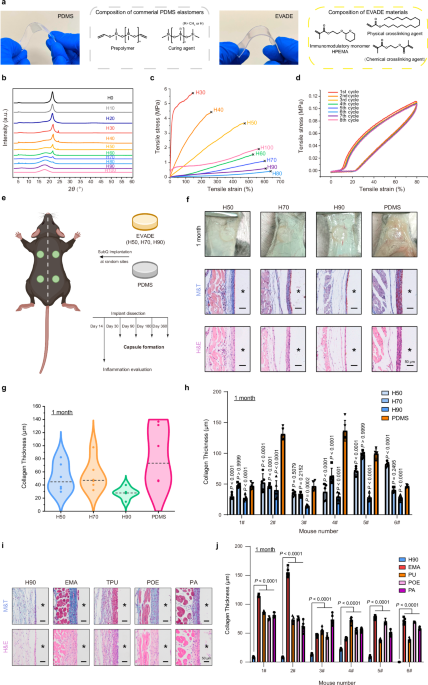
The crystallization behavior of EVADE materials (H0 to H100, 10% as a step) was measured by Powder X-ray Diffractometer (Rigaku Ultima IV) with Cu Kα radiation in the 2θ range 3−60° with a step size of 0.02° (40 mA, 40 kV).
DSC
The Tgs of EVADE materials (H0 to H100, 10% as a step) were measured by DSC (Q20, TA instruments, New Castle, DE, USA) at a scanning range of −70 to 150 °C with a heating/cooling rate of 10 °C min−1 and a nitrogen flow rate of 50 mL min−1. During the test, each sample was heated to 150 °C, kept at 150 °C for 3 min to eliminate thermal history, cooled to −70 °C, and then heated to 150 °C again. Tg was taken as the inflection of the DSC curve.
Mechanical tests
Tensile tests were measured on a tensile-compressive mechanical tester (Zwick/Roell Z020) with a 500 N load cell at room temperature and 37 °C, respectively. For tensile tests, EVADE materials and PDMS were cut from the elastomer sheet into a dumbbell shape with an initial gauge length of 12 mm and a width of 2 mm. The stretch rate was 10 mm min−1, and the nominal stress and strain were recorded. Young’s modulus of the elastomer was calculated from the initial slope of the tensile stress-strain curve with a strain below 10%. Average data were acquired by testing three specimens for each sample. The stress-strain curve of the EVADE elastomer (H90) in cyclic stress-strain tests (80% strain) in successive stretching was conducted using the same size samples as mentioned above, but the stretching speed was set as 5 mm min−1. The authors would like to thank Meifang Wang from Shiyanjia Lab (www.shiyanjia.com) for providing the assistance with the mechanical tests.
Preparation and modulus adjustment of PDMS
PDMS elastomers were prepared and used as controls, with the ratio of prepolymer and curing agent adjusted to ensure a similar tensile modulus between PDMS and H90. The mixture was degassed and poured into plastic dishes at a fixed volume, resulting in a PDMS layer with a thickness of ~1 mm. The PDMS layer was cured at 80 °C for 2 h, peeled, and subsequently cleaned with ethanol and dried.
Scanning electron microscope (SEM)
The elastomer samples were dried thoroughly under vacuum and then loaded on aluminum pellets using carbon tape. The samples were coated with a 2-nm gold layer using a sputter coating system, and images were recorded on a scanning electron microscope (Hitachi SU8010).
Non-contact profilometry
Samples with a diameter of 4 mm were cleaned with 99% isopropyl alcohol. On an anti-vibration table, the samples were placed under the ×20 lens of a non-contact profilometer (KEYENCE VR-5200). The samples were adjusted to ensure a 90° angle between the surface of interest and the lens, and images were focused. Smooth surfaces were acquired in automatic mode; however, if the topography was unknown, the upper and lower z-axis limits were manually adjusted. Using the confocal program visualization mode, the upper and lower z limits were set on the observed image with both focusing extremes appearing completely black. Next, program visualization was set to microscope mode, and brightness was adjusted to automatic, ensuring no areas appeared red due to overexposure. Images and measurements were then captured for each sample. Data were subsequently analyzed.
Biocompatibility of EVADE materials
Human primary umbilical vein endothelial cells (ECs) were isolated from human umbilical veins and arteries with approval from the local medical ethics committee. ECs were cultured in endothelial cell medium supplemented with 10% fetal bovine serum (FBS), 20 mg/mL endothelial cell growth supplement, 60 μg/mL penicillin, and 100 μg/mL streptomycin, maintained at 37 °C in a humidified atmosphere with 5% CO2.
Extraction cytotoxicity test
To assess cytotoxicity, endothelial cell medium extractions were prepared by immersing 600 mg of EVADE material in 3 mL of endothelial cell medium containing 10% FBS for 30 days. Cytotoxicity was evaluated using the Cell Counting Kit-8 (CCK-8) from Meilunbio (catalog no. MA0218). Endothelial cells were seeded at a density of 5000 cells per well in a 96-well plate and incubated overnight. The following day, the culture medium was replaced with the prepared extraction medium. After 24 h of incubation, the medium was replaced with 200 μL of fresh medium containing 20 μL of CCK-8 and incubated for an additional 4 h. Cell viability was measured using a Bio-Rad microplate reader at 450 nm, with 570 nm as the reference wavelength. Cells cultured in standard endothelial cell medium served as controls. Data were averaged from five independent experiments.
Modulation of the RNA profile in macrophages cultured on EVADE materials ex vivo
Murine RAW264.7 macrophages were cultured in RPMI 1640 medium and cultured in the presence of 10% FBS in a humidified atmosphere with 5% CO2 at 37 °C. EVADE materials and PDMS were cut into the size of a six-well plate well and then treated with 75% ethanol. Discard ethanol and let it evaporate until there is no residue. RAW264.7 cells were seeded in a 96-well plate at a density of 50,000 cells mL−1 and incubated for 48 h. The supernatant was collected for cytokine profiling analysis, and then TRIZOL was added to extract RNA for further RNA-seq tests.
Animal work
Mice experiments comply with the relevant regulations, and all protocols are approved by the Institutional Animal Care and Use Committee, Zhejiang Academy of Medical Sciences (Approval number: ZJCLA-IACUC-20010236). Wild-type (WT) female C57BL/6 mice of 6–8 weeks in age were obtained from the animal center of Zhejiang Academy of Medical Sciences, S100a8-KO mice (Strain NO. T011976) were purchased from GemPharmatech (Nanjing, China). The animals were fed a standard laboratory diet and maintained with a 12-hour light/12-hour dark cycle.
For procedures in mice, the EVADE materials (H50, H70, and H90) and PDMS sheets were cut into discs with a biophysical punch (4 mm in diameter). Elastomers samples were sterilized with 75% ethanol, washed with normal saline, and implanted subcutaneously in C57BL/6 female mice. The implantation procedure was as follows. In brief, mice were anesthetized with 3% isoflurane in oxygen, shaved, and disinfected the skin with iodine. An 8 mm longitudinal incision was made on the dorsal surface using surgical scissors to access the subcutaneous space. Then subcutaneous pockets about 0.5 cm away from the incision were created with blunt forceps to implant the elastomer discs. After implantation, the incisions were closed using 5-0 taper-tipped PGA absorbable sutures. Mice were monitored until recovery from anesthesia and raised for 1 day, 2 weeks, 4 weeks, 3 months, 6 months, or longer, respectively. The mice grew normally without discomfort after the implantation, and no body weight loss was observed during the entire experiment.
NHP experiments comply with the relevant regulations, and all protocols are approved by the Laboratory Animal Ethics Committee of the Second Affiliated Hospital, School of Medicine, Zhejiang University (Approval number: 2023-No.30). Cynomolgus monkeys (male; 2 to 3 years old; 2 to 3.5 kg of body weight) were obtained from Suzhou Xishan Zhongke Laboratory Animal Co. Ltd., certified by the Association for Assessment and Accreditation of Laboratory Animal Care. The animals were fed a standard laboratory diet and maintained with a 12-hour light/12-hour dark cycle.
For procedures in NHPs, the animals were anesthetized via intramuscular injection of ketamine (5 mg/kg; Gutian Pharmaceutical) and midazolam (0.2 mg/kg; Ehwa Phrma) and ventilated with room air on an animal ventilator (Matrx, USA). Animals were kept on circulating warm water-based blankets and covered during the entire procedure for body temperature maintenance. An approximately 1 cm dorsal skin incision was made on the thoracic spine’s left and right lateral sides. Blunt dissection was used to create a subcutaneous pocket ventrally about 5 cm deep. H90 and PDMS discs were placed on the left and right subcutaneous pockets. After implantation, the incisions were closed using 5-0 taper-tipped PGA absorbable sutures. All animals received a single 50,000 U/kg perioperative penicillin G benzathine/ penicillin G procaine (Combi-Pen) injection and subcutaneous once-daily meloxicam (0.2 mg/kg on day 1 and 0.1 mg/kg on days 2 and 3) for postsurgical pain.
Retrieval of tissues and elastomers
After two weeks, one month, three months, six months, or longer, mice were sacrificed, and the elastomer samples and the surrounding tissue were excised and collected. The explanted samples were either fixed in 10% formaldehyde solution (for use in histology) or flash frozen (for RNA or protein analysis).
For NHP SubQ retrievals, animal preparation for live excision procedures was carried out at the same time as implantation (described above). Then, biopsy punches (8 mm) were used to sample the skin and SC space at two-month retrieval time points. After retrieval, sites were closed with 5-0 taper-tipped PGA absorbable sutures.
Immunostaining of tissue sections, microscopy, and quantitative image analysis
Tissue samples fixed in paraformaldehyde were dehydrated through a graded ethanol series and embedded in paraffin. Sections (3–5 μm) were cut using a microtome, deparaffinized in xylene, and rehydrated through sequential ethanol and distilled water baths. Sections were stained with hematoxylin, eosin (for cellularity), and Masson’s trichrome (for collagen) following standard histological protocols. For immunohistochemistry, heat-induced epitope retrieval was performed using 0.1 M citrate buffer (pH 6) at 95 °C for 15 min. Sections were then rehydrated with three washes in PBS containing 0.1% Tween. Staining was conducted using a Diaminobenzidine (DAB) chromogenic reagent kit (DAKO, catalog no. K5007). Sections were blocked with peroxidase blocking solution for 10 min at room temperature and washed three times in PBS with 0.1% Tween. Primary antibodies targeting CCR-7 (rabbit IgG; 1:500; Abcam), TNF-α (goat IgG; 15 µg/mL; R&D Systems), IL-10 (goat IgG; 15 µg/mL; R&D Systems), IL-6 (goat IgG; 15 µg/mL; R&D Systems), S100a8 (rabbit IgG; 1:100; Solarbio), MRP8 + MRP14 (mouse IgG1; 1:100; Abcam), α-SMA (mouse IgG2a; 1:400; Boster Bio), and F4/80 (rabbit IgG; 1:5,000; Abcam) were applied. Secondary antibodies conjugated with horseradish peroxidase (anti-mouse, anti-rabbit, or anti-goat) were used for detection. For immunofluorescence, sections were incubated in 0.1 M citrate buffer (pH 6) at 95 °C for 15 min and then rehydrated with three washes in TBS containing 0.025% Triton X-100. After incubation with 10% goat serum for 60 min, sections were exposed overnight to primary antibodies against F4/80 (rabbit IgG; 1:5,000; Abcam), α-SMA (Cy3 polymer; mouse IgG2a; 3 μg/mL; Sigma), CD3 (Brilliant Violet 421 polymer; rat IgG2b; 3 μg/mL; BioLegend), and F4/80 (Alexa Fluor 647 polymer; rat IgG2a; 5 μg/mL; BioLegend). Detection was achieved using Alexa Fluor® 488-conjugated goat anti-rabbit IgG (H + L) F(ab’)2 Fragment (1:200; Cell Signaling Technology) and nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI; 1:50; Sigma–Aldrich). Images were captured with a Nikon intensilight CHGFI microscope and NIS-Elements AR software, and analyzed using a Virtual Slide Microscope (VS120-S6-W, Olympus). Capsule thickness was defined as collagen-rich tissue between the implant surface and adjacent structures. Collagen thickness was quantified from Masson’s trichrome-stained sections. Positive staining areas for TNF-α, IL-6, CCR-7, IL-10, S100a8, α-SMA, and F4/80 were calculated as percentages using ImageJ. All images were processed with Adobe Photoshop 2023, with consistent adjustments to contrast and brightness for representative images.
Protein extraction
Proteins were extracted from skin tissue surrounding H90 implants, PDMS, and non-implanted tissue for further analysis. RIPA lysis buffer (containing phenylmethylsulfonyl fluoride) from Solarbio (catalog no. R0010) was added to the isolated tissue. The tissue was homogenized and lysed on ice for 30 min. Following lysis, the sample was centrifuged at 10,000 g for 5 min at 4 °C, and the supernatant was collected. Protein concentration was determined using a BCA kit from Beyotime (catalog no. P0010S). The protein extract was aliquoted and stored at −80 °C for subsequent use.
Cytokine profiling analysis
A proteome profiler antibody array from the R&D System (catalog no. ARY006) was used for cytokine profiling. After two weeks of implantation, tissue samples were excised and collected, using a mock group without implantation as control. Proteins from the above tissue samples were extracted. Next, 2 mL of blocking buffer was pipetted into each well of the 4-well multi-dish containing each antibody array membrane and incubated for 1 h on a rocking platform shaker. The blocking buffer was aspirated, and 1.5 mL sample solutions containing 300 μg proteins and 15 μL detection antibody cocktail were pipetted to each well and incubated overnight at 2–8 °C on a shaker. Each membrane was carefully placed into individual plastic containers and washed three times with a wash buffer. The membrane was put into the 4-well multidish, containing 2 mL diluted Streptavidin-HRP (1:2,000), and incubated for 30 min on a shaker. After washing three times by wash buffer, the membranes were incubated with 1 mL of Chemi Reagent Mix for 1 min and placed in an autoradiography film cassette (ChemiScope 6000, Clinx) for chemiluminescence. The gray value was measured using ImageJ software.
scRNA-seq analysis
PDMS and H90 discs were implanted into the subcutaneous space of the mice. After one week, implants were removed and washed with sterile PBS to isolate cells from the surface. The resulting cells were pooled per condition and sorted on a Sony MA900 Cell Sorter to filter out dead cells. Libraries were generated from the remaining cells on the DNBelab C4 system according to the manufacturer’s instructions. The barcode sequencing libraries were quantified by Qubit (Invitrogen). The sequencing libraries were sequenced using the DIPSEQ T1 sequencer at the China National GeneBank. The R package Seurat V4 (Satija Lab) was used to cluster the transcripts using K-means clustering and generate UMAP coordinates, visualized on the Loupe Browser 6.0. Clusters were identified as immune cell populations by expression of common immune cell markers, Fn1&Csf1r (macrophages), T/NK cells (Cd3d&Prf1), and dendritic cells (Flt3).
RNA-seq and data analysis
Total RNA was extracted from skin tissue near the implants using TRIzol reagent and treated with an RNase-Free DNase Set (Qiagen). Quantitative RT-PCR was performed with Power SYBR Green Master Mix (Applied Biosystems) following the manufacturer’s guidelines. Extracted RNA (500 ng) was reverse transcribed into complementary DNA (cDNA) using SuperScript™ III First-Strand Synthesis SuperMix (Thermo Fisher). qRT-PCR was conducted on a Real-Time PCR Detection System (CFX384, Bio-Rad, USA) using recommended settings for quantitative and relative expression. Primer sequences used for qRT-PCR are detailed in Supplementary Table 3.
Western blotting
Protein was extracted from skin tissue near the implants by using T-PER Tissue Protein Extraction Reagent (catalog no. 78510, Thermo Pierce) with protease inhibitors (Halt Protease inhibitor single-use cocktail, catalog no. 78445, Thermo Pierce) on ice, and then lysed by sonication (for 30 s on, 30 s off, twice at 70% amplitude). Samples were then subjected to constant agitation for 2 h at 4 °C. Lysates were then centrifuged for 20 min at 7200 g at 4 °C, and protein-containing supernatants were collected in fresh tubes kept on ice. In samples from fat tissue, an excess of fat (a top layer on the supernatant) was first removed before supernatant transfer. 20 µg protein (quantified by BCA assay, BCA protein assay kit, catalog no. P0010, Beyotime) for each lane was boiled at 95 °C for 5 min and electrophoresed on SDS-polyacrylamide gels (Any kD 15-well comb mini-gel, Biorad, catalog no. 456-9036) and then blotted onto nitrocellulose membranes (Millipore, catalog no. IPVH00010). Blots were probed with anti-S100a8 antibody (1:500 dilution; rabbit polyclonal to S100a8; catalog no. 15792-1; Proteintech), anti-IL-17 antibody (1:500 dilution; mouse monoclonal to IL-17; catalog no. 66148-1; Proteintech), anti-IL-1β antibody (1:1,000 dilution; rabbit monoclonal to IL-1β; catalog no. ab254360; Abcam), and anti-TNF-α antibody (1:1,000 dilution; rabbit polyclonal to TNF-α; catalog no. ab6671; Abcam) and anti-β-actin antibody (1:4,000 dilution; monoclonal anti-β-actin antibody produced in mouse; catalog no. A1978; Sigma Aldrich), and anti-GAPDH antibody (1:10,000 dilution; monoclonal anti-GAPDH antibody produced in rabbit; catalog no. ab181602; Abcam) as loading controls followed by goat anti-mouse (1:5000 dilution; catalog no. 62-6520; Thermo Fisher) and goat anti-rabbit (1:10,000 dilution; catalog no. 31460; Thermo Fisher) HRP-conjugated secondary antibodies. Antibody-antigen complexes were visualized using SuperSignal™ West Dura (Thermo Fisher, catalog no. 34075).
Quantitative PCR assay
Total RNA was isolated from skin tissue adjacent to the implants using TRIzol reagent and an RNase-Free DNase Set (Qiagen). Complementary DNA (cDNA) was synthesized from 500 ng of RNA with a SuperScript™ III First-Strand Synthesis SuperMix (Thermo Fisher). Quantitative real-time PCR (qRT-PCR) was performed with Power SYBR Green Master Mix (Applied Biosystems) following the manufacturer’s guidelines. qRT-PCR was carried out on a Real-Time PCR Detection System (CFX384, Bio-Rad, USA), using the recommended settings for quantitative and relative expression analysis. The primers used are detailed in Supplementary Table 3.
Interleukin-17 (IL-17) and glyceraldehyde-3-phosphate dehydrogenase (GAPDH)’s relative levels tested by enzyme-linked immunosorbent assay (ELISA)
An IL-17 ELISA kit from Absin (catalog no. 520009) was used to test the concentration of IL-17 in extracted protein samples. After the implantation of elastomers for one month, tissue samples from implantation were excised and collected. The tissue protein was extracted using a mock group without implantation as a control. In brief, 35 μg extracted protein solutions were added to 96-well plates pre-coated with capture antibody and incubated for 2 h, followed by three washes with wash buffer. The detection antibody was then added to plates and incubated for 2 h, followed by three washes with wash buffer. Next, the streptavidin-HRP was added to plates and incubated for 20 min, followed by three washes with washing buffer. Finally, the 3, 3’, 5, 5’-tetramethylbenzidine (TMB) chromogen solution was added to the plates. After 20 min incubation, the enzymatic reaction was stopped by adding an equal volume of 2 N H2SO4. Absorbance values at 450 nm and 540 nm (as calibration wavelength) were recorded by a microplate reader (MODEL 550, Bio-Rad). Average data were acquired from five specimens.
At the same time, the concentration of GAPDH was measured to correct the experimental results. After the implantation of elastomers for one month, tissue samples from implantation were excised and collected, using a mock group without implantation as the control. In brief, the 96-well plate was incubated with a capture antibody from Beyotime (dilution 1:3,000; catalog no. AG019) overnight at 4 °C, followed by three washes with wash buffer. 10 μg extracted protein samples were added to the plate and incubated for 2 h, followed by 3 washes with wash buffer. Afterward, the plate was incubated with HRP-conjugated detection antibody from Proteintech (dilution 1:50,000; catalog no. 60004) for 2 h, followed by 3 washes with wash buffer. Finally, a TMB chromogen solution was added to the plates. After 20 min incubation, the enzymatic reaction was stopped by adding an equal volume of 2 N H2SO4. Absorbance values at 450 nm and 540 nm (as calibration wavelength) were recorded by a microplate reader (MODEL 550, Bio-Rad). Average data were acquired from five specimens.
Surface protein analysis of elastomers
The in vitro fibrinogen adsorption and in vivo protein adsorption 24-hour post-implantation of EVADE and PDMS elastomers were investigated.
Fibrinogen fouling test by enzyme-linked immunosorbent assay (ELISA)
Fibrinogen (Fg) was selected as the test protein to examine the anti-fouling property of the EVADE elastomer samples (H50, H70, and H90). In brief, EVADE materials were cut from the 1 mm thick elastomer sheet into 4 mm diameter discs with a biopuncher. In addition, a 1 mm thick PDMS sheet and 2-methacyloyloxyethyl phosphorylcholine (MPC) hydrogel sheet were cut into 4 mm diameter discs with a biopuncher and used as controls. To make MPC hydrogels, 200 mg MPC and 20 mg MBAA were dissolved in 1 mL distilled water followed by the addition of 4 mg APS and 4 μL TEMED. After the vortex, the solution was immediately injected into the mold and reacted for 24 h. The resulting hydrogels were soaked in PBS (pH 7.4) for 48 h to remove the residual small molecules. The samples were first placed in a 24-well plate and incubated in PBS for 5 min to equilibrate. Next, the samples were incubated with 1 mL of 1 mg mL−1 Fg in PBS buffer for 1 h, followed by 3 washes with PBS buffer. Samples were then transferred to new wells and incubated with 1 mL of horseradish peroxidase (HRP) conjugated anti-fibrinogen (1 μg mL−1) from Bellancom Chemistry (catalog no. G11390) in PBS buffer for 0.5 h. All samples were then transferred to new wells after another five washes with pure PBS buffer. Afterward, 1 mL of OPD chromogen solution was added. After 15 min incubation, the enzymatic reaction was stopped by adding an equal volume of 1 N HCl. The absorbance value at 490 nm was recorded by a microplate reader (MODEL 550, Bio-Rad) and was normalized to that of the PDMS sample. Average data were acquired from six specimens.
Proteomics
After the elastomers were implanted subcutaneously for 24 h, remove them from the body, peel off the visible attached tissue, gently rinse with PBS, and store at −80 °C. For this assay, proteins adsorbed on the elastomer surface were reduced (5 mM dithiothreitol; 56 °C for 30 min) and alkylated (11 mM iodoacetamide; room temperature in the dark for 15 min). Trypsin was added at 1:50 trypsin-to-protein mass ratio for the first digestion overnight and 1:100 trypsin-to-protein mass ratio for a second 4 h-digestion. The digested peptides were desalted by a Strata X C18 column (Phenomenex, Torrance, CA, USA) and lyophilized. The peptides were labeled based on the manufacturing instructions of the TMTsixplex (Thermo Fisher Scientific). Tryptic peptides were separated by high-pH reversed-phase high-performance liquid chromatography (HPLC) using an Agilent 300Extend C18 column (5 μm particle size, 4.6 mm inner diameter, 250 mm length; Agilent, Fremont, CA, USA). Peptide samples were analyzed using an EASY-nLC 1000 LC system (Thermo Fisher Scientific, Waltham, MA, USA) coupled with an Orbitrap Fusion mass spectrometer (Thermo Fisher Scientific) and subsequently analyzed on a Q Exactive Plus mass spectrometer (Thermo Fisher Scientific). MS/MS files were searched against the UniProt human proteome database (20380 sequences) using MaxQuant (version 1.5.2.8) software enabled with the Andromeda search engine. The quantification analysis was performed at the protein level by MaxQuant software. Peptides were desalted using C18 SpinTips (Protea) then lyophilized and stored at −80 °C. Per the manufacturer’s instructions, peptides were labeled with TMTsixplex (Thermo Fisher Scientific). Peptides were then loaded on a pre-column and separated by reverse-phase high-performance liquid chromatography (Easy nLC 1000; Thermo Fisher Scientific) over a 140-min gradient before nanoelectrospray using a Q Exactive mass spectrometer (Thermo Fisher Scientific). Raw mass spectral data files were searched using Proteome Discoverer (Thermo Fisher Scientific) and Mascot version 2.4.1 (Matrix Science). Tandem mass tag (TMT) quantification was obtained using Proteome Discoverer and isotopically corrected per the manufacturer’s instructions.
Mice models of inhibiting and knocking out of S100A8/S100A9
Two mice models were constructed to confirm the relationship between S100A8/A9 expression and the FBR.
Inhibition of S100A9 with taquimod
For mice implanted with PDMS, the distribution of tasquinimod at 1 mg/kg/day and 3 mg/kg/day (administered orally via drinking water) started the day after implantation and continued for one month.
Generation of S100A8 conditional knockout (cKO) mice
In brief, S100a8 conditional knockout homozygote (cHO) mice (Lyz2-iCre+/−, S100a8fl/fl) were produced by crossing S100a8 conditional knockout heterozygote (cHE) mice (Lyz2-iCre+/−, S100a8fl/+) with homozygous S100a8 floxed mice (Lyz2-iCre− /−, S100a8fl/fl). Both S100a8-CKO (Strain NO. T011059) and Lyz2-iCre (Strain NO. T003822) were purchased from GemPharmatech (Nanjing, China). S100a8 cHO mice were generated through two rounds of crossing: homozygous S100a8-CKO mouse (S100a8fl/fl) ×Cre-positive mouse (Lyz2-iCre+/−), and then crossing with the resulting heterozygous S100a8 cHE mice (Lyz2-iCre+/−, S100a8fl/+).
Diabetes induction in mice
Wild-type (WT) male C57BL/6 mice, aged 6–8 weeks, were sourced from the Zhejiang Academy of Medical Sciences animal center. To induce diabetes, mice were given daily intraperitoneal injections of 50 mg/kg body weight STZ (10 mg/mL in citrate buffer, pH 4.5) for five consecutive days while maintaining a non-fasted regular diet. Fourteen days post-injection, body weight and blood glucose levels were assessed to confirm diabetes. Mice with non-fasted blood glucose levels exceeding 19.4 mmol/L for two consecutive days were deemed diabetic.
Fabrication of EVADE CSII catheter
To prepare the EVADE CSII catheter, a stainless steel wire with a diameter of 400 μm was immersed in sucrose solution and dried to form a soluble layer, facilitating easy demolding. Subsequently, the wire was soaked in a reaction mixture containing HPEMA monomer, ODA monomer, and 0.5 wt% (relative to the weight of monomers) of photoinitiator Irgacure 819. The stainless steel surface was then dip-coated with this mixture to form a uniform polymer solution layer at a speed of 10 mm/min. Polymerization occurred under UV irradiation (365 nm) at approximately 180 mW/cm2 for 10 min. After polymerization, the elastomer samples were immersed in a sterile PBS buffer (pH 7.4) for two days, demolding and drying under vacuum at room temperature. Subsequently, the catheter was cut from the infusion set, and adhesive was used to affix the EVADE CSII catheter onto the infusion set.
In vivo insulin infusion catheter implantation in mice
Diabetic C57BL/6 mice were shaved and anesthetized using 3% isoflurane in oxygen, shaved, and disinfected the skin with iodine. A longitudinal incision of about 4 mm was made on the dorsal surface using surgical scissors to provide access to the subcutaneous space. Then subcutaneous pockets about 0.5 cm away from the incision were created with blunt forceps to implant the catheters. After implantation, the incisions were closed using 5-0 taper-tipped PGA absorbable sutures. Mice were monitored until recovery from anesthesia and raised for one month. The mice grew normally without discomfort after the implantation, and no body weight loss was observed during the entire experiment.
Insulin pump therapy in mice
Diabetic mice with newly induced diabetes had their dorsal skin shaved and disinfected with iodine. An infusion catheter was then inserted subcutaneously. To evaluate the performance of EVADE CSII catheters under natural micromotion, the catheter was connected to an infusion line and the mouse was allowed to wear the complete infusion set, including the catheter and line, for up to 7 days without restriction in a single cage. For assessing continuous insulin infusion via EVADE CSII catheters, the infusion set was connected to an insulin pump (Medtronic MiniMed 722) filled with 0.2 IU/mL insulin. Mice received a continuous basal insulin infusion at a rate of 10 μL/h, with three additional bolus insulin infusions of 45 μL each. On day seven post-catheter insertion, an insulin absorption assay was performed by administering a bolus infusion of 2 IU/kg insulin through the pump. Blood glucose levels were measured at predefined intervals using a glucometer following insulin infusion. Serum insulin levels were determined by centrifuging blood samples (3000 × g, 4 °C) for 15 min, followed by quantification with the Elabscience human insulin ELISA kit (E-EL-H2665). Throughout the study, mice were fasted but had ad libitum access to water. Mice receiving subcutaneous insulin injections at the same dosage served as controls.
Statistics and reproducibility
Details of the sample size and appropriate statistical test are included in the figure captions. The data are expressed as means ± s.d. The data were analyzed for statistical significance by two-side one-way analysis of variance (ANOVA) with Bonferroni multiple comparison correction or unpaired, two-tailed t-test using SPSS Statistics 26.0. No adjustments were made for multiple comparisons. All in vitro and in vivo results are representative of three to six independents.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
link